酵母表面展示技术平台
酵母表面展示(Yeast surface display,YSD)是一种强大的蛋白质工程技术,是目前蛋白工程中重要的高通量筛选平台。酵母细胞具有真核细胞翻译后修饰机制,能够帮助目的蛋白正确折叠,可以用来展示各种真核蛋白,包括抗体、受体、酶和抗原肽等。
阿帕克生物具有经验丰富的专业团队和先进技术平台,结合流式细胞分选提供基于酵母表面展示技术的蛋白质工程服务,用于改善蛋白质性质,包括亲和力、特异性、酶功能和稳定性等,欢迎咨询与合作交流!
技术原理
NBbiolab酵母表面展示平台采用“α-凝集素”展示系统,将纳米抗体基因序列和表达标签HA tag基因序列插入至蛋白质支架-Aga2基因的展示质粒载体中,该载体使用营养标记物在酵母中维持选择性生长,并通过培养基中加入半乳糖诱导酵母进行表面展示,纳米抗体融合蛋白被分泌并锚定在酵母细胞壁上。再结合磁珠及流式分选技术进行多轮筛选,可筛选得到高亲和力、高稳定性的克隆。
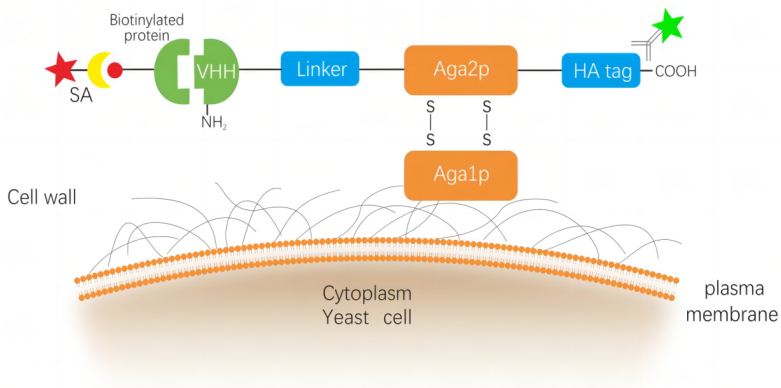
图 阿帕克生物基于a-凝集素的 Saccharomyces cerevisiae 酵母表面展示系统示意图
酵母表面展示技术优势
NBbiolab HITSmart纳米抗体发现服务
阿帕克生物将酵母表面展示、细胞流式分选与NGS结合使用,解放常规文库筛选的局限,实验效率翻倍,一次实验可获得超越传统筛选方式的百倍优质克隆!已为多家企业成功筛选到高亲和力抗体(靶点有HSA等)阻断抗体(靶点有PD-L1、PD-1、CTLA-4、LAG3、TIGIT等),特异性抗体(靶点有CD16a,FcRL5等)和TCR mimic 抗体。
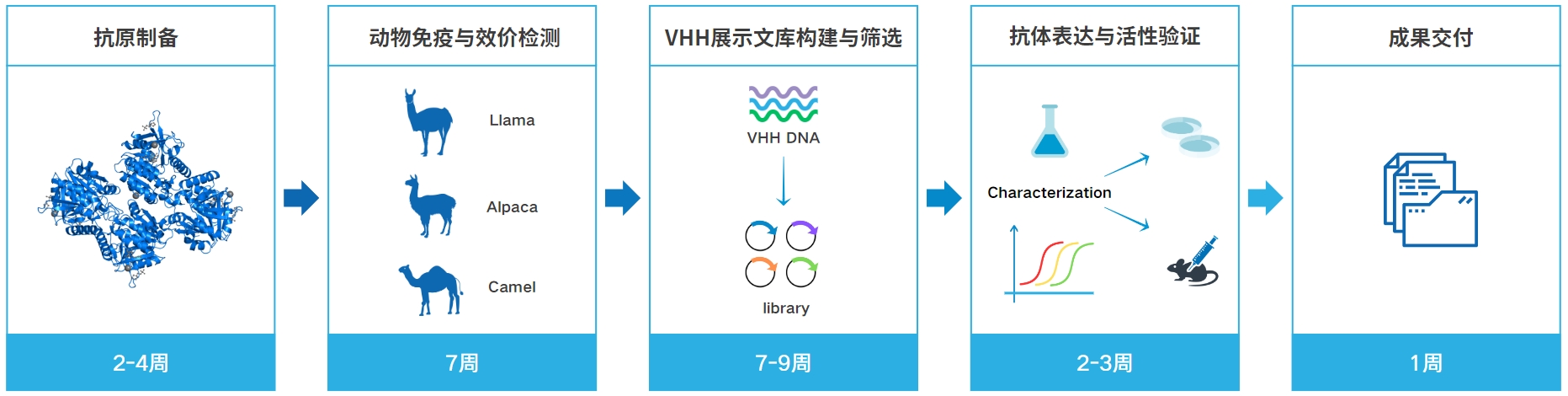
NBbiolab HITSmart纳米抗体服务流程
NBbiolab HITSmart纳米抗体服务优势
【相关阅读】
酵母表面展示VS噬菌体展示(点击篇名跳转阅读)
【摘要】酵母表面展示与噬菌体展示相比,具有显著优势。酵母具有类似于高等真核生物的分泌途径,使蛋白质能够正确地折叠和分泌。与噬菌体表面展示不同的是,使用FACS可以基于亲和力和展示水平筛选库,对于在酵母表面展示中的高亲和力的选择不受表达水平的负面影响,这消除了表达偏向可能引起的任何潜在偏差。此外,酵母表面展示可实时区分仅在抗体亲和力上略有不同的克隆。通过两色染色,可以直接在酵母细胞表面确定抗体亲和力,避免了亚克隆、表达和纯化的需要。这种技术使得在一种展示格式中,可以快速筛选和优化目标抗体的亲和力、稳定性和细胞外表达。
酵母展示技术用于提高蛋白质的亲和力(点击篇名跳转阅读)
【摘要】酵母展示技术可以区分亲和力只有2倍差异的蛋白质,具有较高的敏感性,已经成为蛋白质亲和力成熟的重要工具。如图 4 所示,一般通过随机突变创建大小为107 −109 的蛋白质突变文库,经电转化和同源重组,这些突变体与 Aga2p 融合展示在酵母表面;通过与荧光标记配体一起孵育,用FACS分选高亲和力突变体,每一轮分选的细胞可以扩增培养参与再次分选,从而降低文库多样性,最终获得少量的高亲和力酵母细胞克隆;也可以提取所选酵母细胞 DNA,进行新一轮随机突变,构建新的展示文库,参与新一轮分选,直到筛选出所需亲和力的蛋白质。
酵母展示技术用于TCR工程化改造(点击篇名跳转阅读)
【摘要】体内天然的TCRs虽然表现出与抗体相似的遗传和序列多样性,但TCRs的结合亲和力比抗体要低几个数量级。尽管TCR亲和力与T细胞活性之间的相关性存在一些分歧,但选择高亲和力TCR或通过定向进化微调TCR的亲和力都是改善TCR-T抗肿瘤反应的技术手段。通过酵母表面展示技术对TCR的重链和轻链可变区的互补决定区进行随机突变得到酵母展示文库,经过多轮筛选,定向进化后的TCR可实现皮摩尔范围内的亲和力。与噬菌体展示方法相比,使用酵母表面展示TCR定向进化的一个主要优点是它能够直接评估在酵母细胞上表达的TCR或突变体的结合亲和力,而不需要经过大量的蛋白质表达和纯化步骤。
酵母表面展示技术用于酶的定向进化(点击篇名跳转阅读)
【摘要】酶蛋白有广泛的应用价值,参与各种化学反应,包括分解、合成和能量转化等。
通过定向进化,人们可以调整酶蛋白的结构和活性,使其更具高效性和特异性。使用酵母展示技术已成功实现了酶的人工定向进化。多种酶已成功展示在酵母细胞表面。米黑根毛霉脂肪酶(RML)是一种生物催化剂,通过定点突变并展示在酵母细胞表面,得到的突变体显示出比原始酶活性更高的酯化活性,可应用于食品工业、精细化工和生物柴油生产。 抗坏血酸过氧化物酶(APEX)是一种工程过氧化物酶,通过酵母展示系统进行定向进化,创造了进化的分裂APEX2系统,具有较高的过氧化物酶活性,可应用于新领域的生物学研究。使用酵母展示系统展示的β-葡萄糖苷酶,可以增加饮料的香气和风味。